Biochemistry & Evolution
Nitrogen Fixation & Nitrogenase Biology
Understanding nitrogenase means understanding both where it came from and how its complex machinery is built. My work bridges these two perspectives: tracing the evolutionary origins of nitrogen fixation and dissecting how nitrogenase assembles its metal cofactors and performs catalysis. Together, these approaches reveal why this enzyme looks the way it does, how it functions, and how its complexity might be re-engineered.
Origins & Evolution
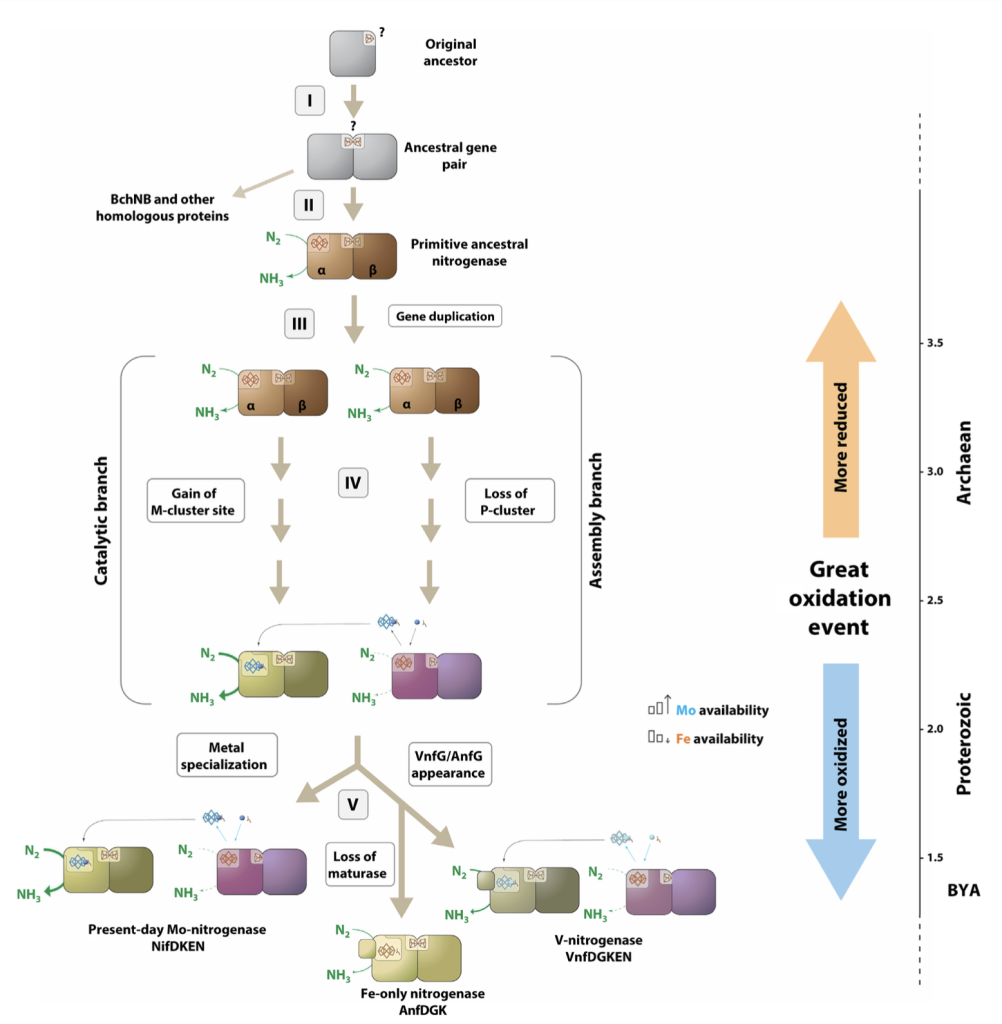
I investigate how nitrogenase emerged, diversified, and adapted across early microbial lineages. By comparing homologs, mapping conserved structural motifs, and examining simplified or ancestral-like variants, I explore why nitrogen fixation adopted its current architecture and metal dependence. This work asks fundamental questions about early Earth, cofactor availability, and the evolutionary pressures that shaped one of biology’s most ancient and consequential enzymes.
Assembly, Cofactors & Mechanism
Modern nitrogenase depends on intricate metalloclusters and a multi-step maturation pathway. I study how these clusters form, how the component proteins coordinate electron flow, and how fully active nitrogenase can be assembled outside its native host. Through heterologous expression, biochemical characterization, and structural analysis, I dissect how the enzyme is built, what limits activity, and how these insights can inform future engineering.
Related publications:
- Y. A Liu, C. C. Lee, K. Górecki, M. T Stiebritz, C. Duffin, J. B. Solomon, M. W. Ribbe, Y. Hu, Heterologous synthesis of a simplified nitrogenase analog in Escherichia coli, (2025), Science Advances, 11(18). Read
- C. C. Lee*, K. Górecki*, M. Stang, M. W. Ribbe, and Y. Hu, Cofactor maturase NifEN: A prototype ancient nitrogenase?, (2024), Science Advances, 10(24). Read
- J. B. Solomon*, Y. A. Liu*, K. Górecki*, R. Quechol, C. C. Lee, A. J. Jasniewski, Y. Hu, M. W. Ribbe, Heterologous expression of a fully active Azotobacter vinelandii nitrogenase Fe protein in Escherichia coli, (2023), mBio, 14:e02572-23. Read
- R. Quechol, J. B. Solomon, Y. A. Liu, C. C. Lee, A. J. Jasniewski, K. Górecki, P. Oyala, B. Hedman, K. O. Hodgson, M. W. Ribbe, and Y. Hu, Heterologous synthesis of the complex homometallic cores of nitrogenase P- and M-clusters in Escherichia coli, (2023), Proceedings of the National Academy of Sciences USA, 120 (44) e2314788120. Read (open access at PubMed)
- M.W. Ribbe, K. Górecki, M. Grosch, J.B. Solomon, R. Quechol, Y.A. Liu, C.C. Lee, and Y. Hu, Nitrogenase Fe Protein: A Multi-Tasking Player in Substrate Reduction and Metallocluster Assembly, (2022), Molecules, 27, 6743. Read
Bacterial Efflux Pumps
Organisms have several ways to defend themselves against toxic compounds. Some destroy the molecule outright, like β-lactamases that break open the β-lactam ring of penicillins. Others survive by altering the toxin’s target so it can no longer bind. But bacteria also rely on a third strategy that is just as important: they actively pump harmful substances out of the cell before they can do damage.
These systems are known as efflux pumps, and the Resistance Nodulation Division (RND) family is one of the most striking examples. RND pumps span the entire cell envelope and form a continuous tunnel that ejects antibiotics, toxins, and even metal ions directly into the outside environment.

What makes these pumps surprising is how many of them bacteria carry. E. coli has five RND systems with almost overlapping functions. Removing one hardly changes anything. Only when all five are deleted does the cell struggle. This raises an interesting question: if these pumps are so redundant, why do some of them have features that have been preserved for billions of years?
This puzzle led us to the MdtABC system. Unlike most RND pumps, which have a single inner membrane component, MdtABC contains two, MdtB and MdtC. Our phylogenetic analysis showed that this duplication happened only once, early in bacterial evolution, and that the overall architecture of the complex has remained unusually stable ever since. Such long-term conservation suggests a specific and irreplaceable function.
What that function is remains unknown, but its evolutionary footprint hints that the MdtABC pump plays a role that other systems cannot fully substitute. Understanding that role is the next step.
Related publications:
- K. Górecki & M.M. McEvoy, Phylogenetic analysis reveals an ancient gene duplication as the origin of the MdtABC efflux pump, (2020), PLOS ONE 15(2): e0228877. Read
Copper Homeostasis
Copper is essential but toxic in excess, and bacteria are especially sensitive to elevated levels. E. coli relies on several systems to manage this stress, including the plasmid-encoded pco operon, long viewed as a periplasmic copper-defense mechanism. Within this system, PcoB sits in the outer membrane, yet its function has been unclear.
Our structural and biochemical work showed that PcoB forms a classical β-barrel with a highly electronegative funnel leading from the extracellular side toward the periplasm. Rather than acting as an efflux channel, this architecture supports Cu²⁺ uptake, consistent with metal-binding data and with the broader roles of PcoA, PcoC, and PcoE in periplasmic copper handling.
During copper stress, bacteria restrict outer membrane permeability, yet the periplasm still requires Cu²⁺ not only for detoxification but also for essential processes such as respiration. Our findings support the idea that PcoB provides a controlled influx pathway, supplying copper to periplasmic systems while preventing uncontrolled accumulation.

(Li et al., Protein Science 2022).
Related publications:
- N. Nayeri, K. Górecki, K. Lindkvist-Petersson, P. Gourdon, P. Li, Isolation and crystallization of copper resistance protein B (CopB) from Acinetobacter baumannii, (2025), Protein Expression and Purification, 227, 106635. Read
- N. Nayeri, P. Li, K. Górecki, K. Lindkvist-Petersson, P. Gourdon, Principles to recover copper-conducting CTR proteins for the purpose of structural and functional studies, (2023), Protein Expression and Purification, 203, 106213. Read
- P. Li, N. Nayeri, K. Górecki, E. Ramos Becares, K. Wang, D. Ram Mahato, M. Andersson, S.Abeyrathna, K. Lindkvist-Petersson, G. Meloni, J. Winkel Missel, P. Gourdon, PcoB is a defense outer membrane protein that facilitates cellular uptake of copper, (2022), Protein Science 31(7), e4364. Read (published earlier as a preprint on bioRxiv: Read)
Respiratory Chain Complex I & Sodium Homeostasis
Respiratory complex I is one of the central engines of cellular energy conversion, transferring electrons to quinone while pumping ions across the membrane to power ATP synthesis. Its membrane arm is built from the NuoL, NuoM, and NuoN subunits, long suspected to have evolved from ancient Na⁺/H⁺ antiporters. This idea has been supported by structural and sequence similarities, but functional evidence has been limited.
In our work, we tested this evolutionary link directly by expressing the complex I subunits in Bacillus subtilis strains lacking the core components of the Mrp antiporter system. The imported NuoLMN proteins were able to replace MrpA or MrpD and restore ion-transport activity, demonstrating that these membrane proteins remain functionally compatible despite billions of years of divergence. This result shows that complex I’s membrane arm is built on an ancestral antiporter architecture that continues to shape its mechanism today.
Related publications:
- E. Sperling, K. Górecki, T. Drakenberg, C. Hägerhäll, Functional Differentiation of Antiporter-Like Polypeptides in Complex I; a Site-Directed Mutagenesis Study of Residues Conserved in MrpA and NuoL but Not in MrpD, NuoM, and NuoN, (2016), PLOS ONE 11, 7. Read
- K. Górecki, C. Hägerhäll, T. Drakenberg, The Na+ transport in Gram-positive bacteria defect in the Mrp antiporter complex measured with 23Na-NMR, (2014) Analytical Biochemistry 445: 80-86. Read (open access at PubMed)
- V. K. Moparthi, B. Kumar, Y. Al-Eryani, E. Sperling, K. Górecki, T. Drakenberg, C. Hägerhäll, Functional role of the MrpA- and MrpD-homologous protein subunits in enzyme complexes evolutionary related to respiratory chain complex I, (2014), BBA – Bioenergetics 1837: 178-185. Read
- E. Virzintiene, V. K. Moparthi, Y. Al-Eryani, L. T. Shumbe, K. Górecki, C. Hägerhäll, Structure and function of the C-terminal end of MrpA in the Bacillus subtilis Mrp-antiporter complex – the evolutionary progenitor of the long, membrane parallel helix in Complex I. (2013) FEBS Letters 587 (20): 3341-3347. Read
- T. Gustavsson, M. Trane, V. K. Moparthi, E. Miklovyte, L. Moparthi, K. Górecki, T. Leiding, S. Petersson Årsköld, C. Hägerhäll, A cytochrome c-fusion protein domain for convenient detection, quantification and enhanced production of membrane proteins in Escherichia coli – expression and characterisation of cytochrome-tagged complex I subunits, (2010) Protein Science 19: 1445-1460. Read